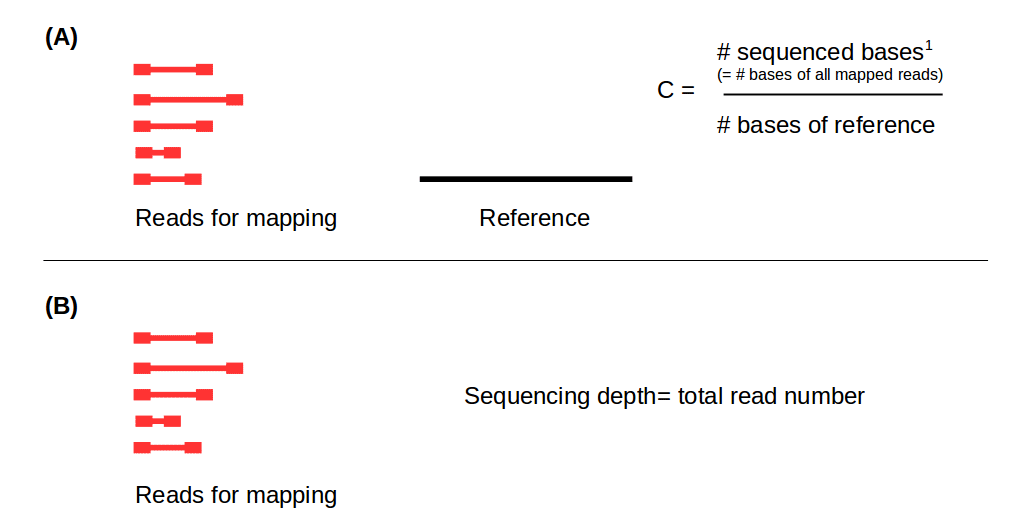
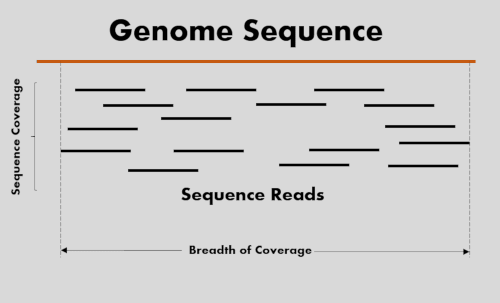
GitHub - GenomicaMicrob/coverage_calculator: A simple script to calculate the coverage of a genome assembly

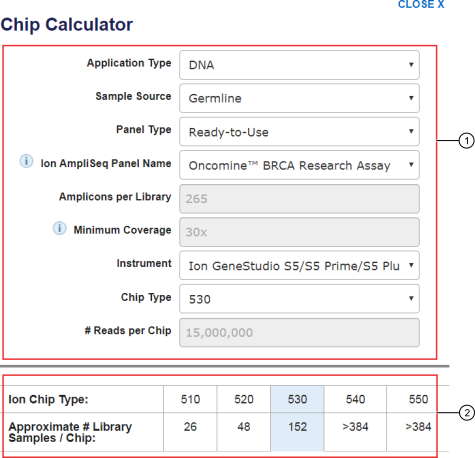
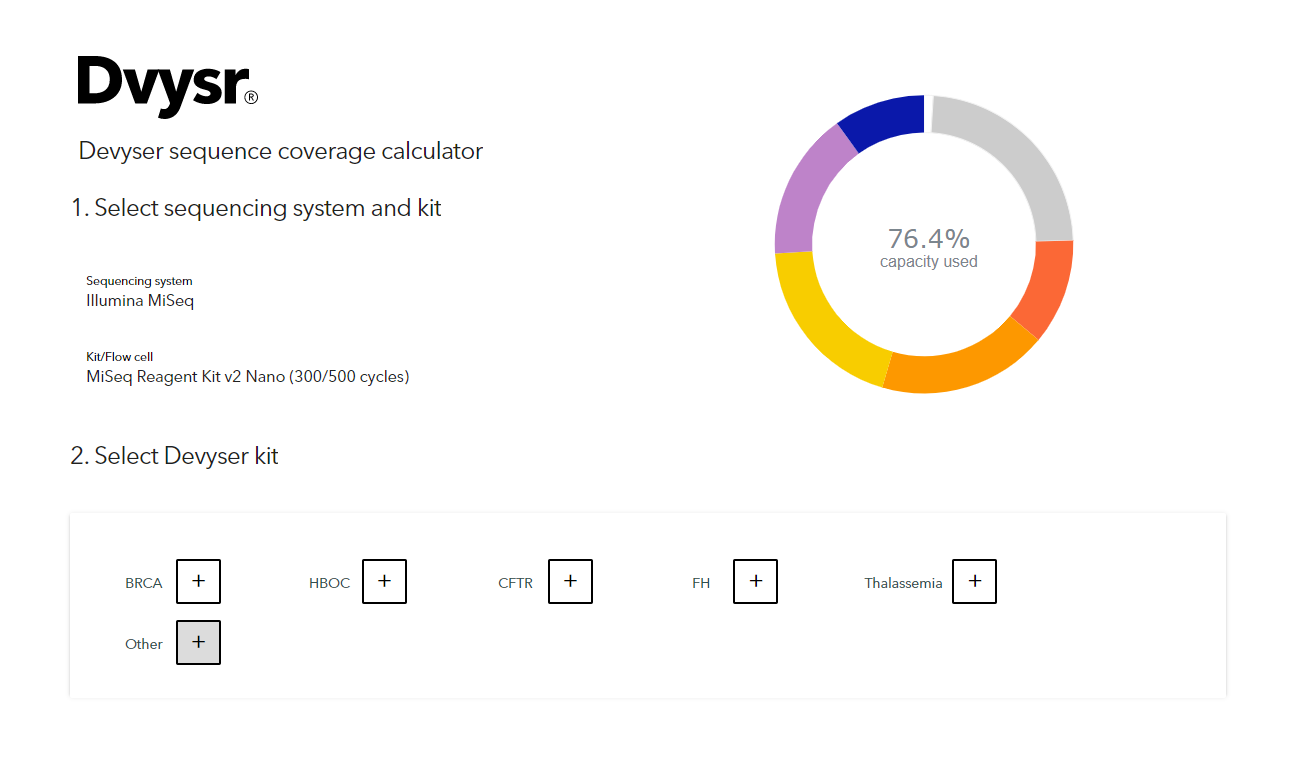
Devyser on X: "Did you know we have a Coverage Calculator which can help your sequencing planning? Just select your system and kit, the number and type of samples, and easily calculate

Covcalc: Shiny App for Calculating Coverage Depth or Read Counts for Sequencing Experiments | R-bloggers